CodeMol
Desktop protein visualization meets terminal-style power
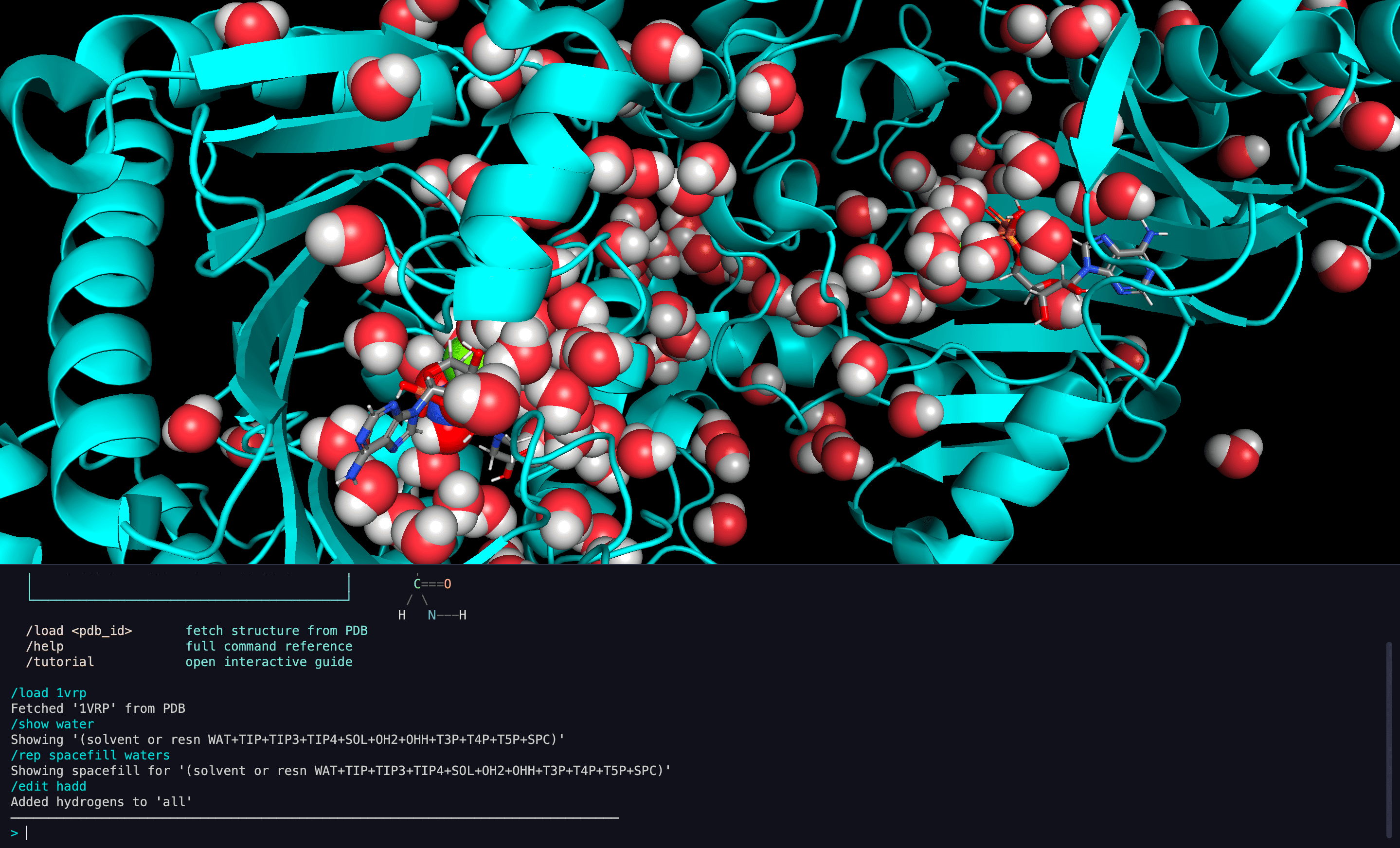
CodeMol is a desktop protein visualization application that replaces traditional GUI menus with a terminal-style command console. Every molecular operation — from loading structures to running analyses — is driven by text commands, giving researchers the speed and composability of a command line with the visual power of a 3D molecular viewer.
Under the hood, CodeMol provides 200+ discoverable Python tools organized across 27+ groups. Each tool follows a consistent run() interface, making the system infinitely extensible — from basic visualization presets to complex docking workflows.
With AI agent integration, researchers can describe tasks in natural language. Specialist LLM agents route requests to the appropriate tools, whether it's setting up a docking simulation or analyzing a molecular dynamics trajectory.
The application provides:
- •A terminal-style command console for all molecular operations, replacing traditional menus with composable text commands.
- •200+ Python tools across 27+ groups, each following a consistent run() interface for visualization, analysis, and manipulation.
- •AI agent integration with specialist routing — describe tasks in natural language and let LLM agents compose the right tool sequences.
- •Real-time collaborative sessions via WebSocket relay, enabling shared molecular analysis across multiple viewers.
- •Trajectory analysis with MD playback, RMSD tracking, clustering, and multi-structure comparison workflows.
CodeMol is designed for researchers who think in commands, not clicks.
By combining a 3D rendering engine with a programmable console, it gives computational biologists the flexibility to build custom workflows without leaving the viewer.
A molecular viewer built for power users.
Features
Console-driven molecular visualization with AI-powered workflows
Console-First Interaction
Terminal-style command console replaces traditional GUI menus. Text commands for all molecular operations with tab completion and command history.
200+ Discoverable Tools
Organized across 27+ groups covering visualization, selection, measurement, structural analysis, file I/O, and more — all accessible via a run() interface.
AI Agent Integration
Natural language interaction via LLM agents with specialist routing for docking and trajectory analysis. Describe what you want in plain English.
Multi-Structure Workflows
Load and compare multiple molecular structures simultaneously. Overlay, align, and analyze differences across models with automatic command scoping.
Collaborative Sessions
Real-time collaboration via WebSocket relay. Hosts broadcast actions to multiple viewers for shared molecular analysis sessions.
Trajectory Analysis
Molecular dynamics playback with frame controls, RMSD tracking, hydrogen bond monitoring, clustering analysis, and multi-structure comparison.
CodeMol in Action

Main Viewer Interface
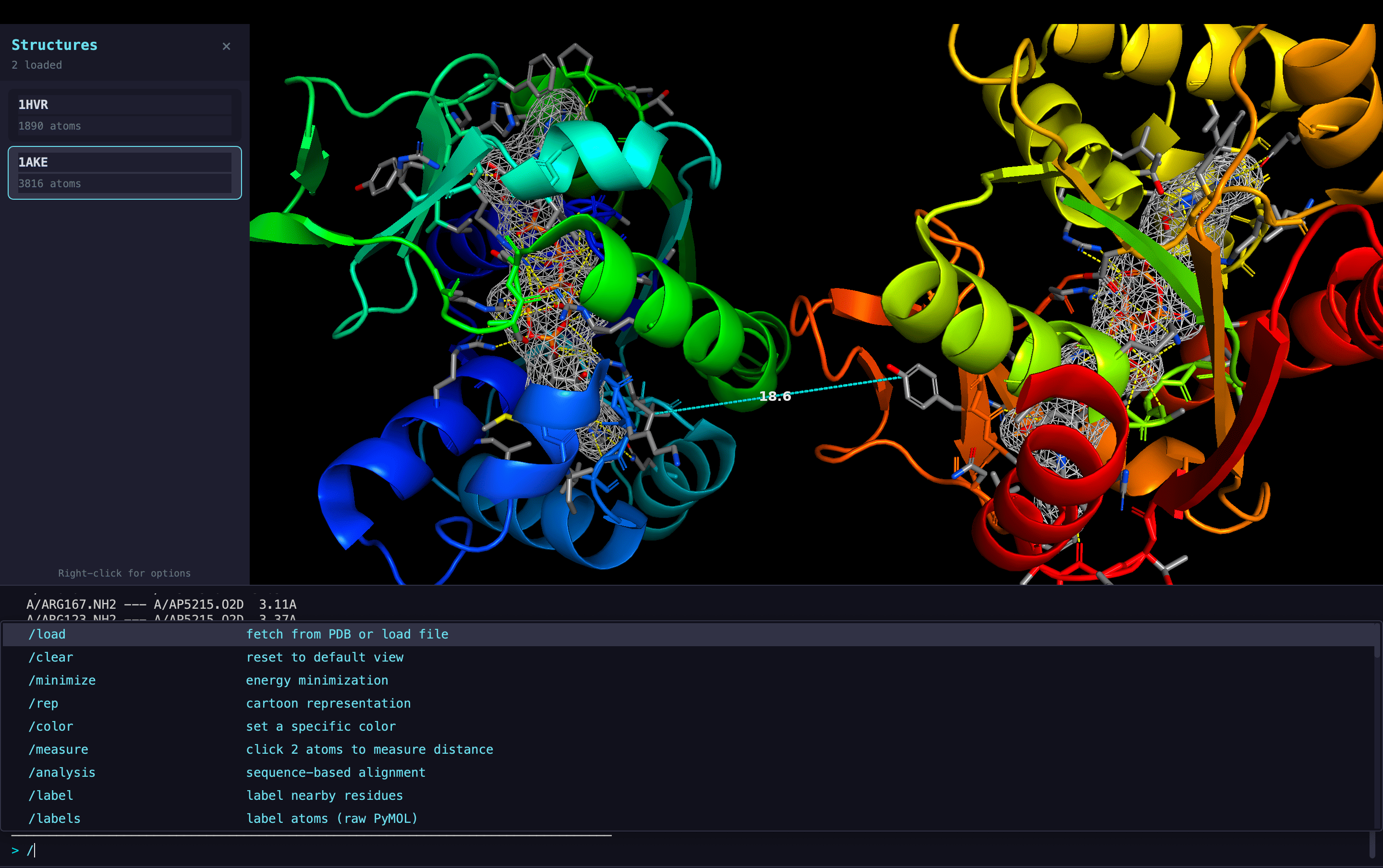
Molecular Representations
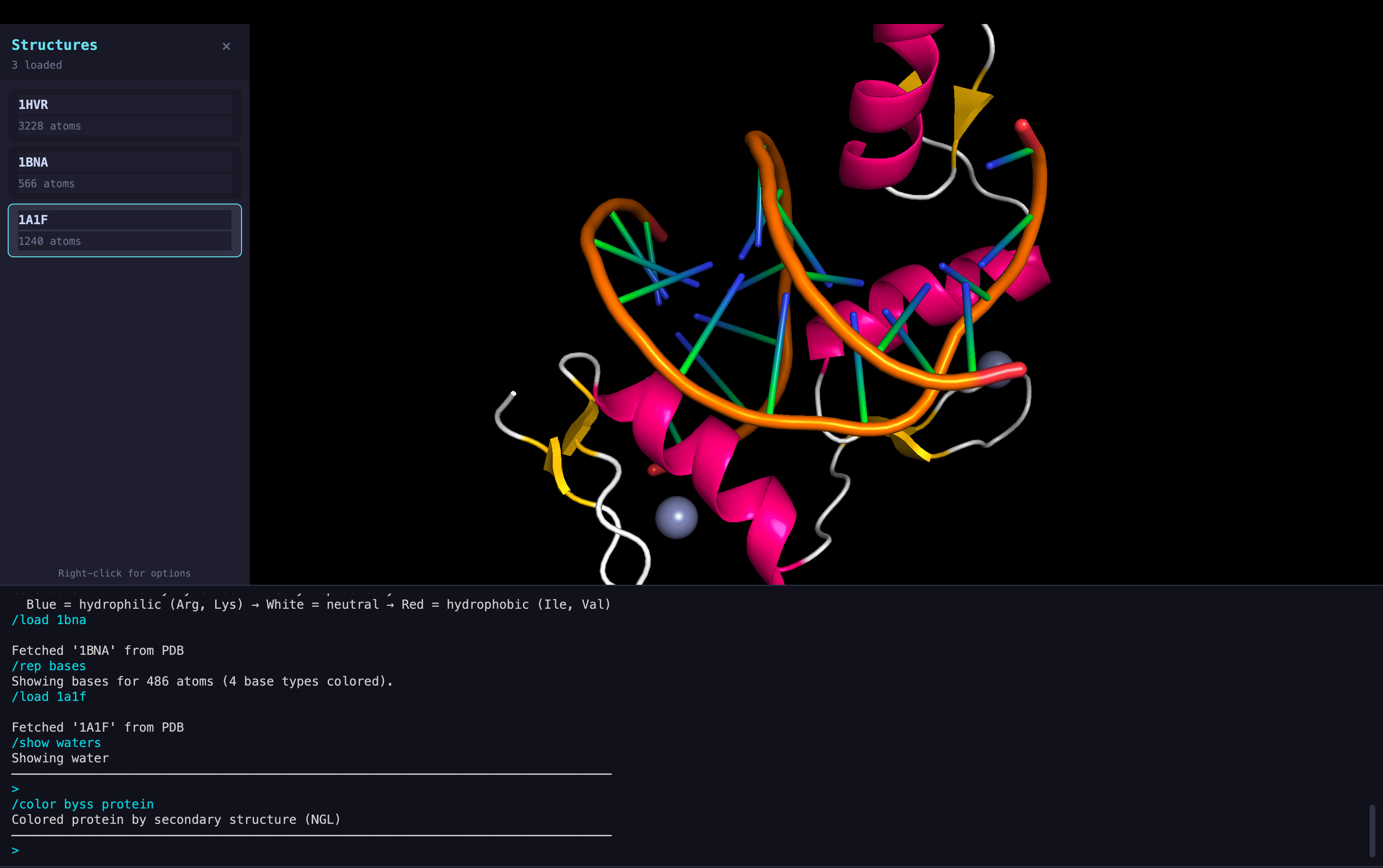
DNA Visualization
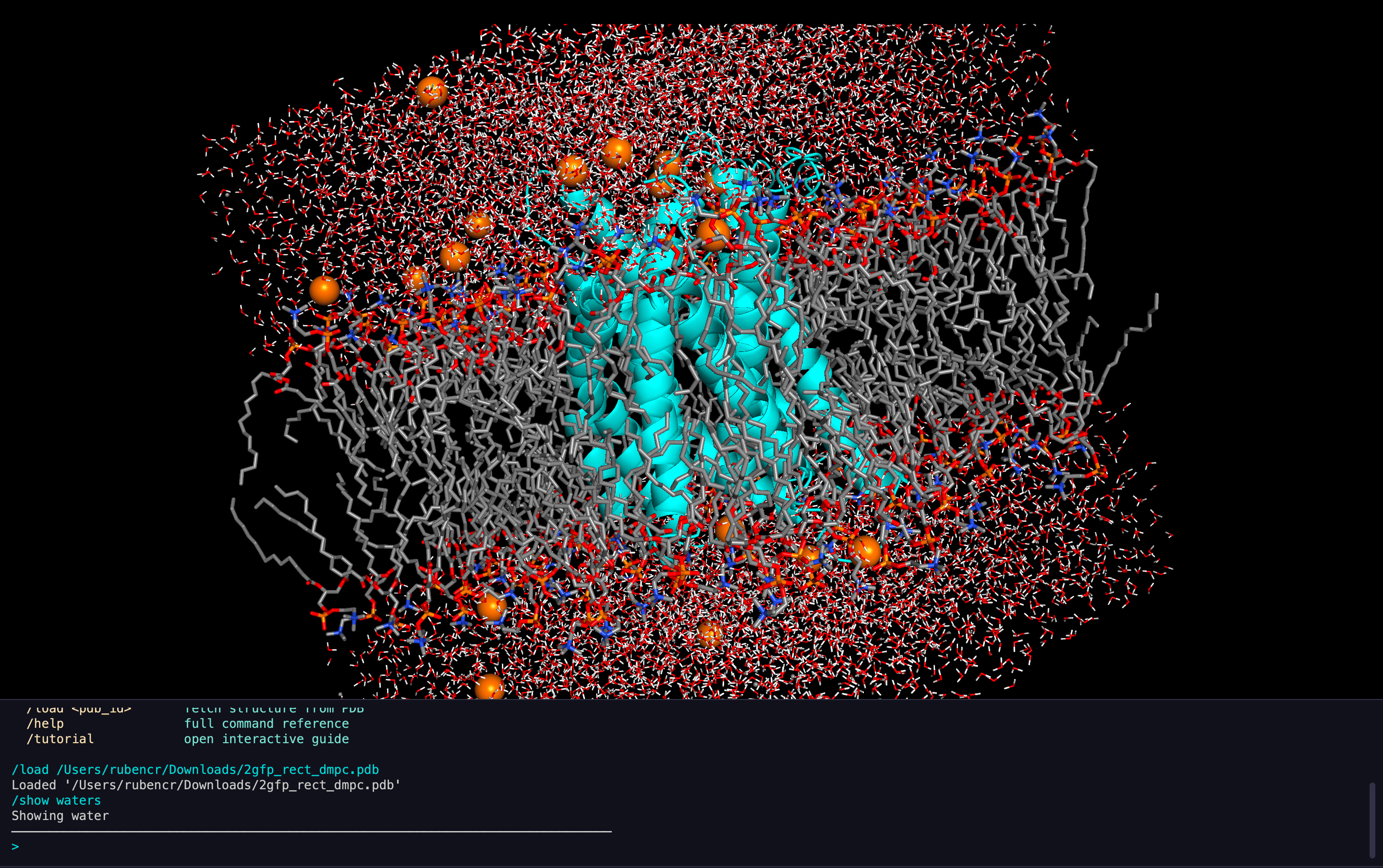
Membrane Systems
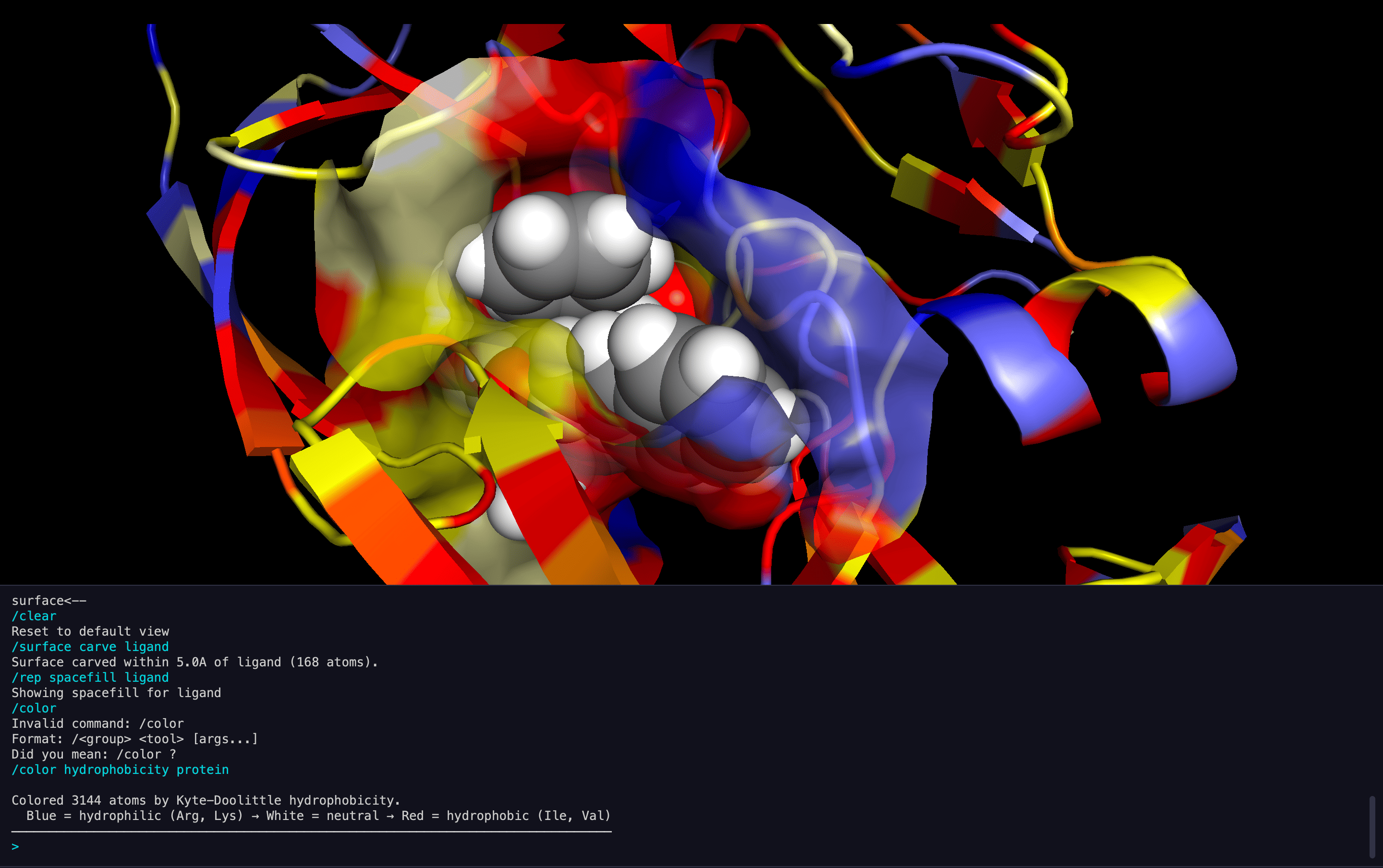
Cavity Detection
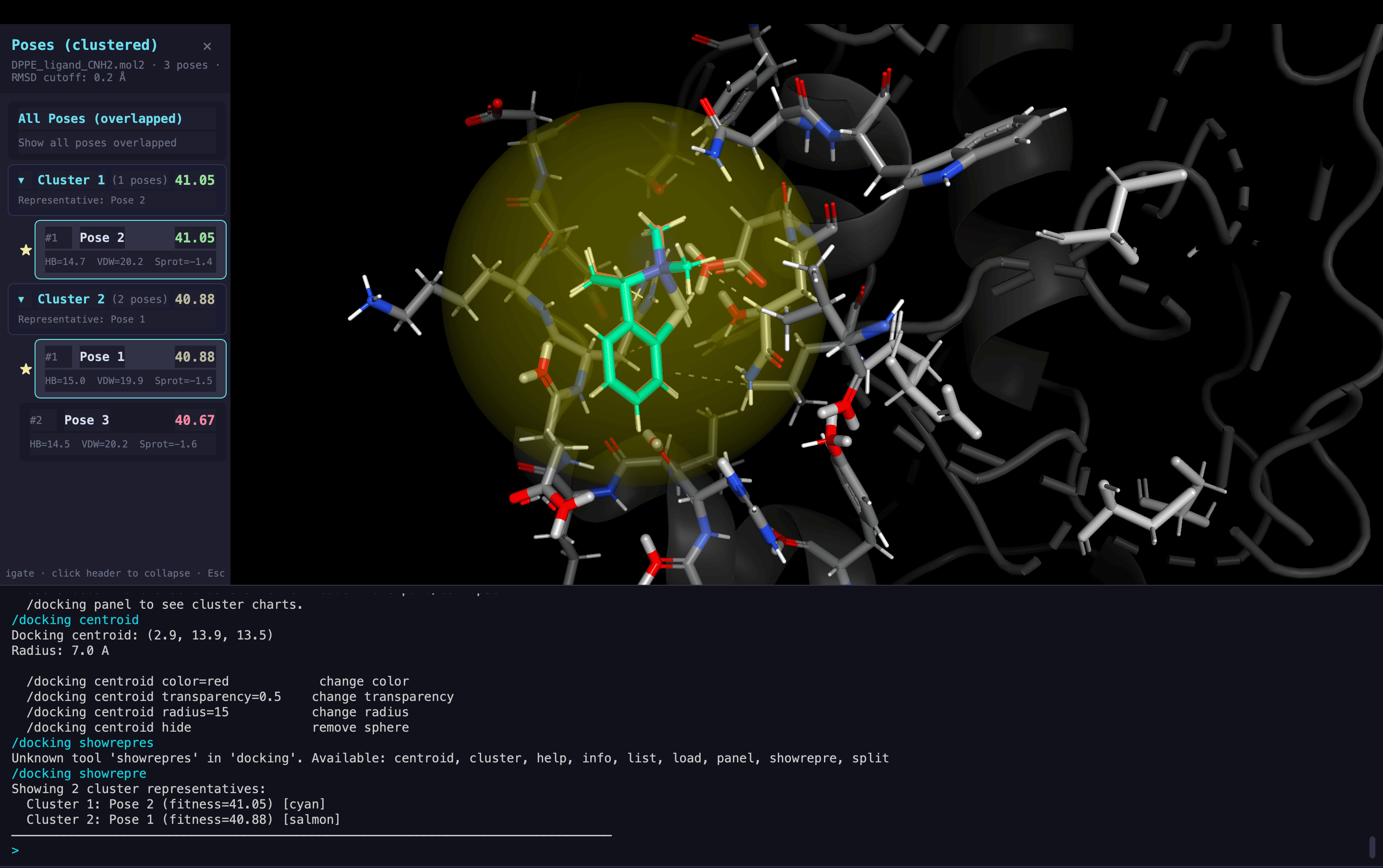
Molecular Docking
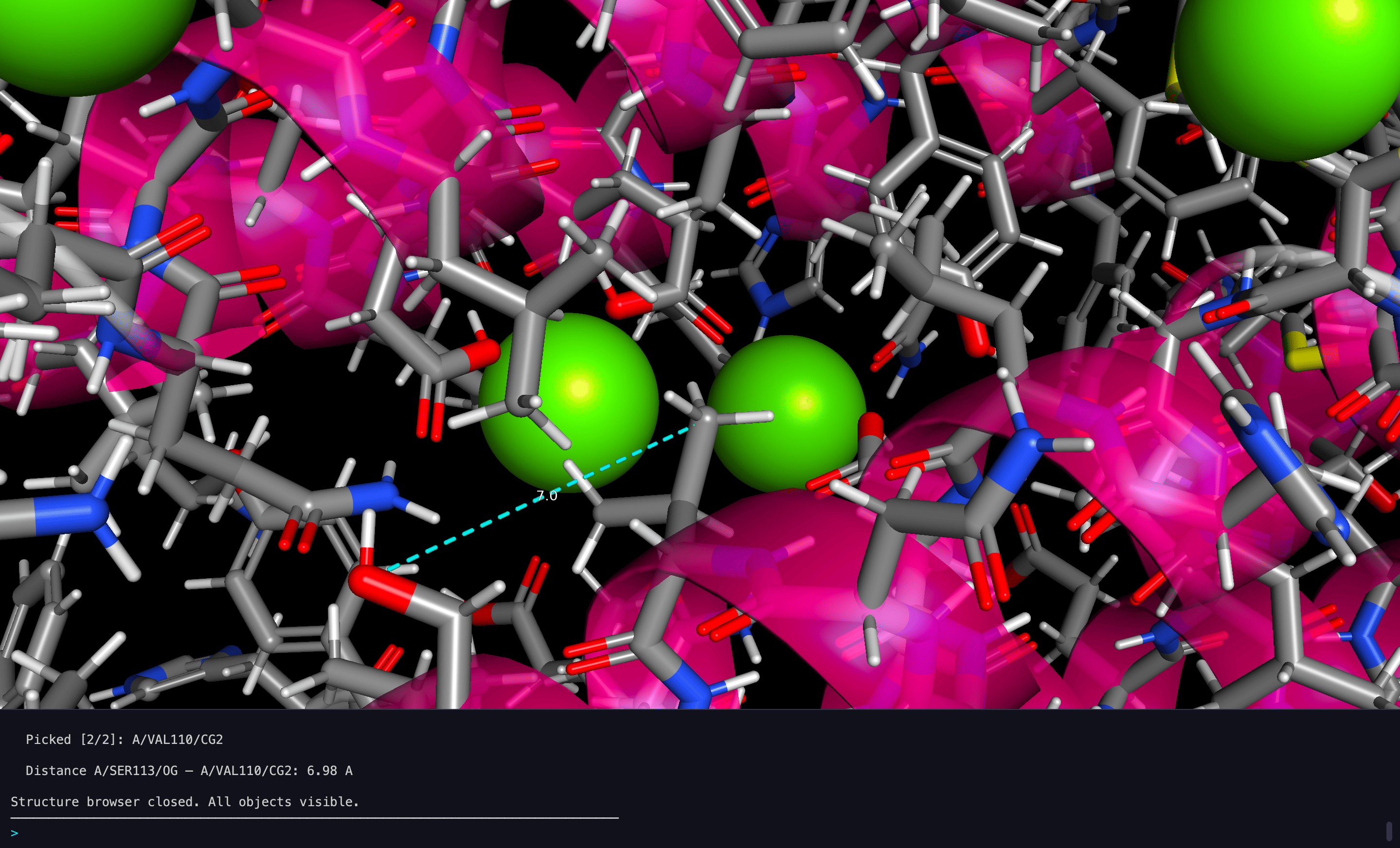
Ion Visualization
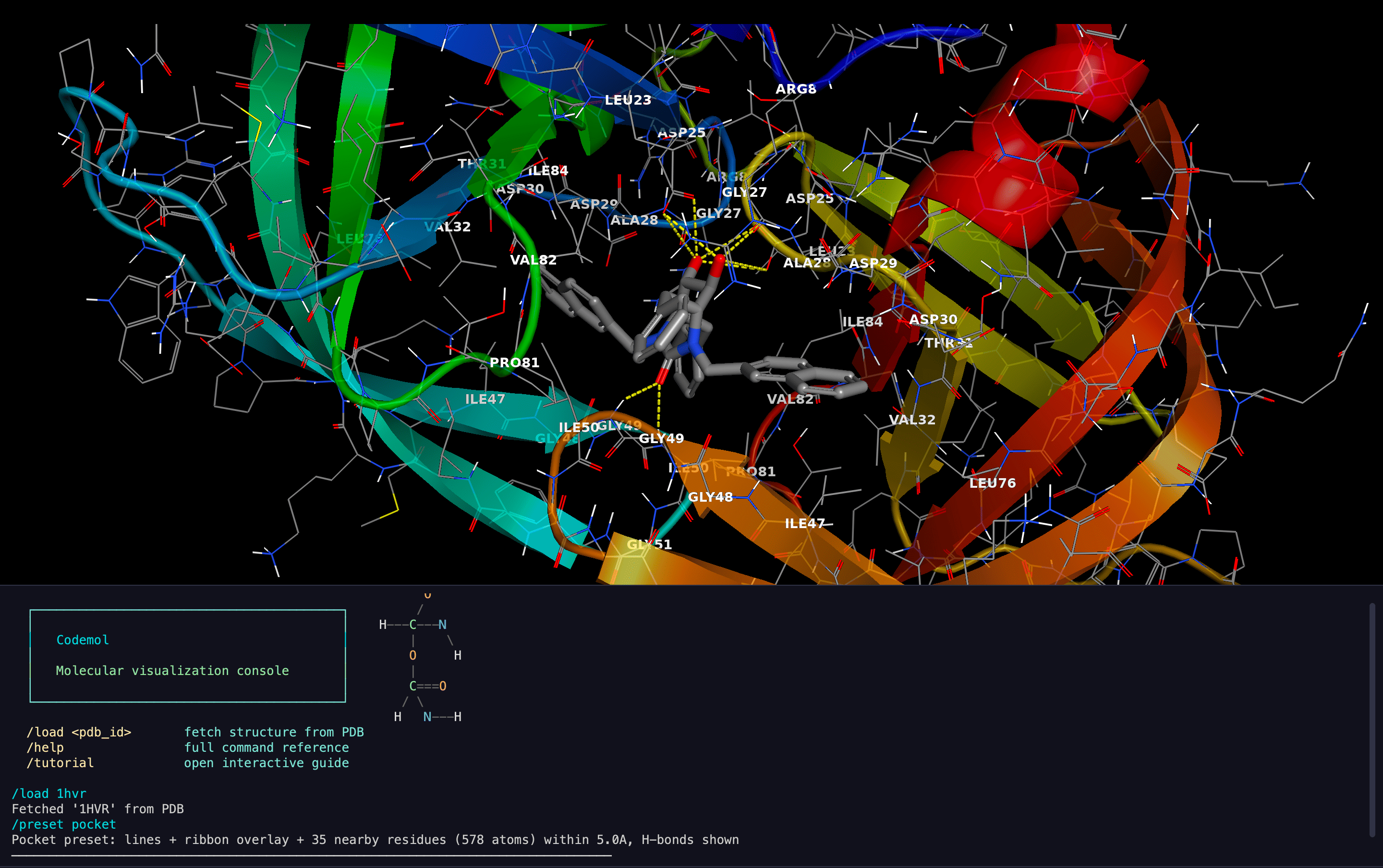
Visual Presets
Protein Structure